

| Pathway Id | ko04122 |
| Class & Sub-Class | Class: Genetic Information Processing Sub-Class: Folding sorting and degradation |
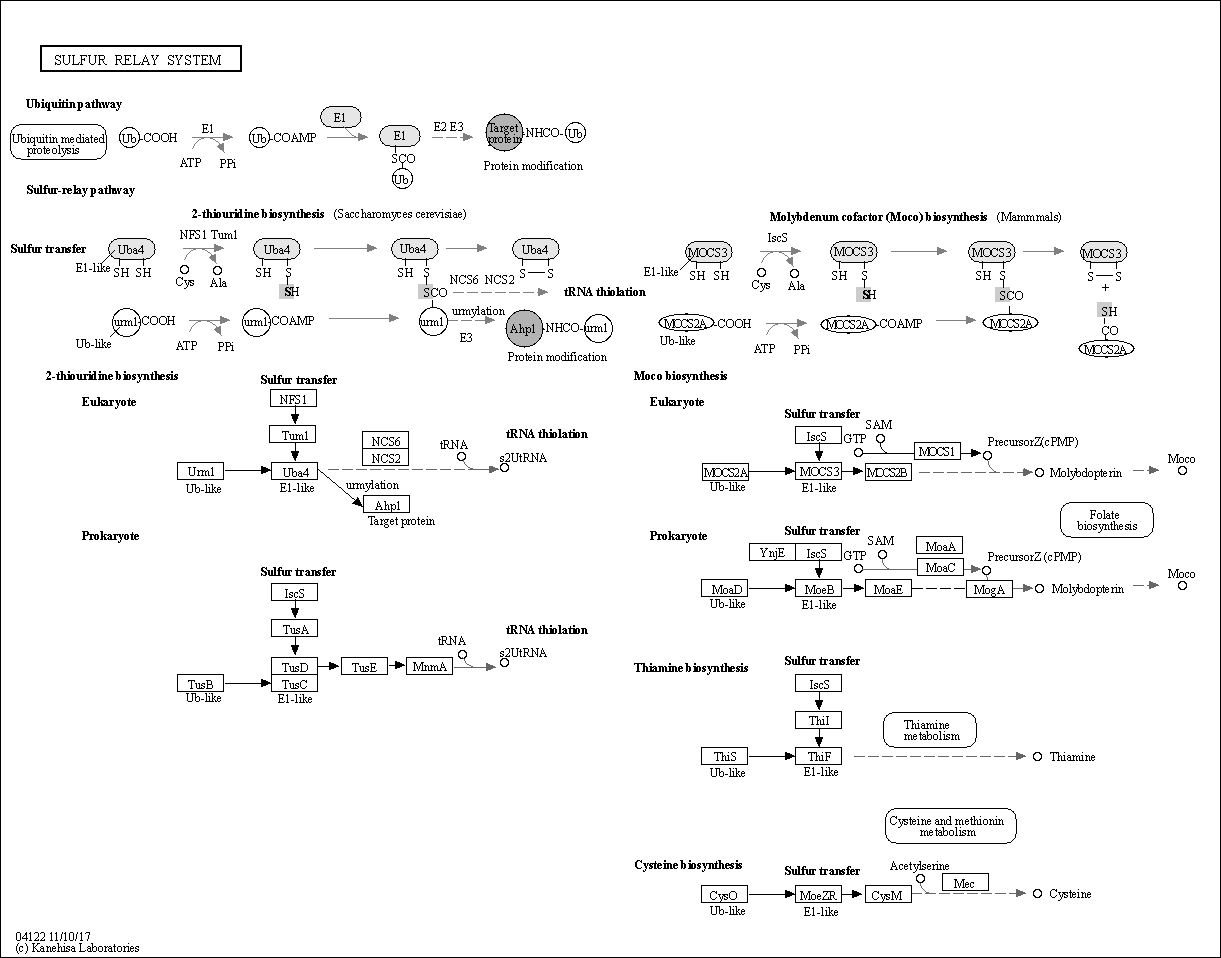
| Description | Ubiquitin and ubiquitin-like proteins (Ubls) are signalling messengers that control many cellular functions; such as cell proliferation; apoptosis; and DNA repair. It is suggested that Ub-protein modification evolved from prokaryotic sulfurtransfer systems. Molybdenum cofactor (Moco) and thiamin are sulfur-containing cofactors whose biosynthesis includes a key sulfur transfer step that uses unique sulfur carrier proteins; MoaD and ThiS. Ubiquitin; MoaD; and ThiS are all structurally related proteins whose C-termini are activated through adenylation by homologous E1-like enzymes. s2T biosynthesis may share similar chemistry with Moco and thiamin synthesis. In Saccharomyces cerevisiae; Urm1 and Uba4 function as part of a ubl protein conjugation system; though they have sequence homology to bacterial sulfur-transfer enzymes and the ability to function in sulfur transfer. Source: KEGG Database |
Make changes to the option below (image Size) and click here to update the map:
| Image Size | You can change the size of an image just by changing the width or height attributes. Size (enter width only) the present size is: | |
| Get Data Out | |
| Click here to show selected items: | |