

| Pathway Id | ko03030 |
| Class & Sub-Class | Class: Genetic Information Processing Sub-Class: Replication and repair |
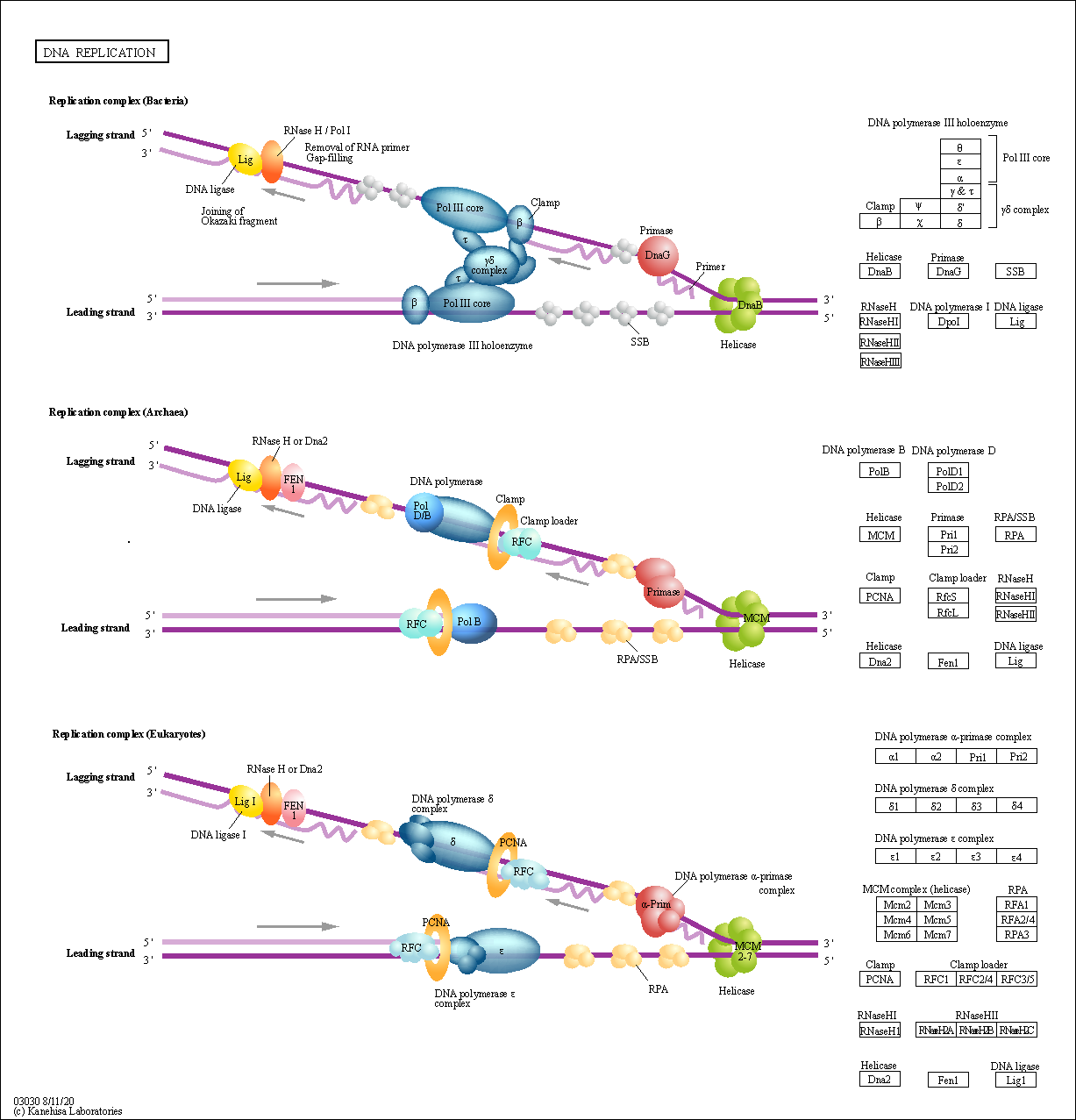
| Description | A complex network of interacting proteins and enzymes is required for DNA replication. Generally; DNA replication follows a multistep enzymatic pathway. At the DNA replication fork; a DNA helicase (DnaB or MCM complex) precedes the DNA synthetic machinery and unwinds the duplex parental DNA in cooperation with the SSB or RPA. On the leading strand; replication occurs continuously in a 5 to 3 direction; whereas on the lagging strand; DNA replication occurs discontinuously by synthesis and joining of short Okazaki fragments. In prokaryotes; the leading strand replication apparatus consists of a DNA polymerase (pol III core); a sliding clamp (beta); and a clamp loader (gamma delta complex). The DNA primase (DnaG) is needed to form RNA primers. Normally; during replication of the lagging-strand DNA template; an RNA primer is removed either by an RNase H or by the 5 to 3 exonuclease activity of DNA pol I; and the DNA ligase joins the Okazaki fragments. In eukaryotes; three DNA polymerases (alpha; delta; and epsilon) have been identified. DNA primase forms a permanent complex with DNA polymerase alpha. PCNA and RFC function as a clamp and a clamp loader. FEN 1 and RNase H1 remove the RNA from the Okazaki fragments and DNA ligase I joins the DNA. Source: KEGG Database |
Make changes to the option below (image Size) and click here to update the map:
| Image Size | You can change the size of an image just by changing the width or height attributes. Size (enter width only) the present size is: | |
| Get Data Out | |
| Click here to show selected items: | |